|
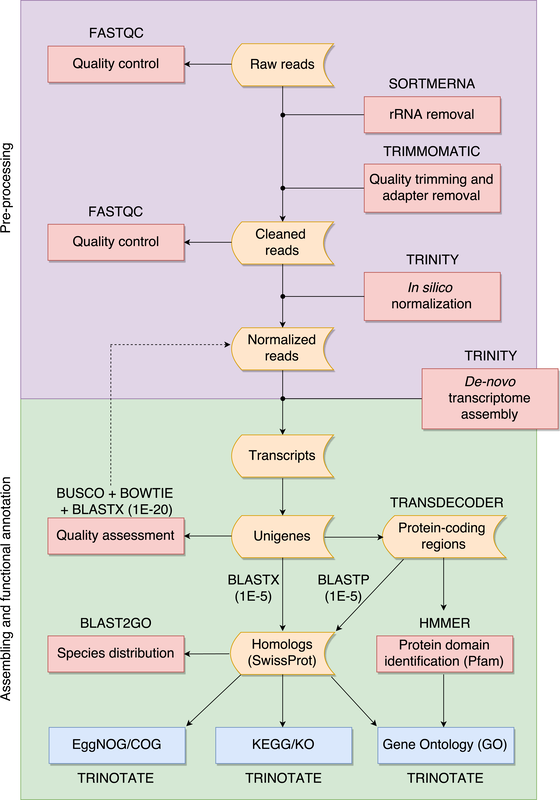
Whole genome sequencing (WGS) is a very valuable resource to understand the evo- lutionary history of poorly known species. However, in organisms with large genomes, as most amphibians, WGS is still excessively challenging and transcriptome sequencing (RNA-seq) represents a cost-effective tool to explore genome-wide variability. Non- model organisms do not usually have a reference genome and the transcriptome must be assembled de-novo. We used RNA-seq to obtain the transcriptomic profile for Oreobates cruralis, a poorly known South American direct-developing frog. In total, 550,871 transcripts were assembled, corresponding to 422,999 putative genes. Of those, we identified 23,500, 37,349, 38,120 and 45,885 genes present in the Pfam, EggNOG, KEGG and GO databases, respectively. Interestingly, our results suggested that genes related to immune system and defense mechanisms are abundant in the transcriptome of O. cruralis. We also present a pipeline to assist with pre-processing, assembling, evaluating and functionally annotating a de-novo transcriptome from RNA-seq data of non-model organisms. Our pipeline guides the inexperienced user in an intuitive way through all the necessary steps to build de-novo transcriptome assemblies using readily available software and is freely available at:
https://github.com/biomendi/TRANSCRIPTOME- ASSEMBLY-PIPELINE/wiki Full reference: Montero-Mendieta S, Grabherr M, Lantz H, De la Riva I, Leonard JA, Webster MT, Vilà C. (2017) A practical guide to build de-novo assemblies for single tissues of non-model organisms: the example of a Neotropical frog. PeerJ 5, e3702 [PDF]
0 Comments
Leave a Reply. |
Hi, I am Santi. This blog series was mainly created to include a summary of each of my publications. However, this blog is also a place where I will write about science and my life as researcher in the field of evolutionary biology.
Categories
All
Powered by
|